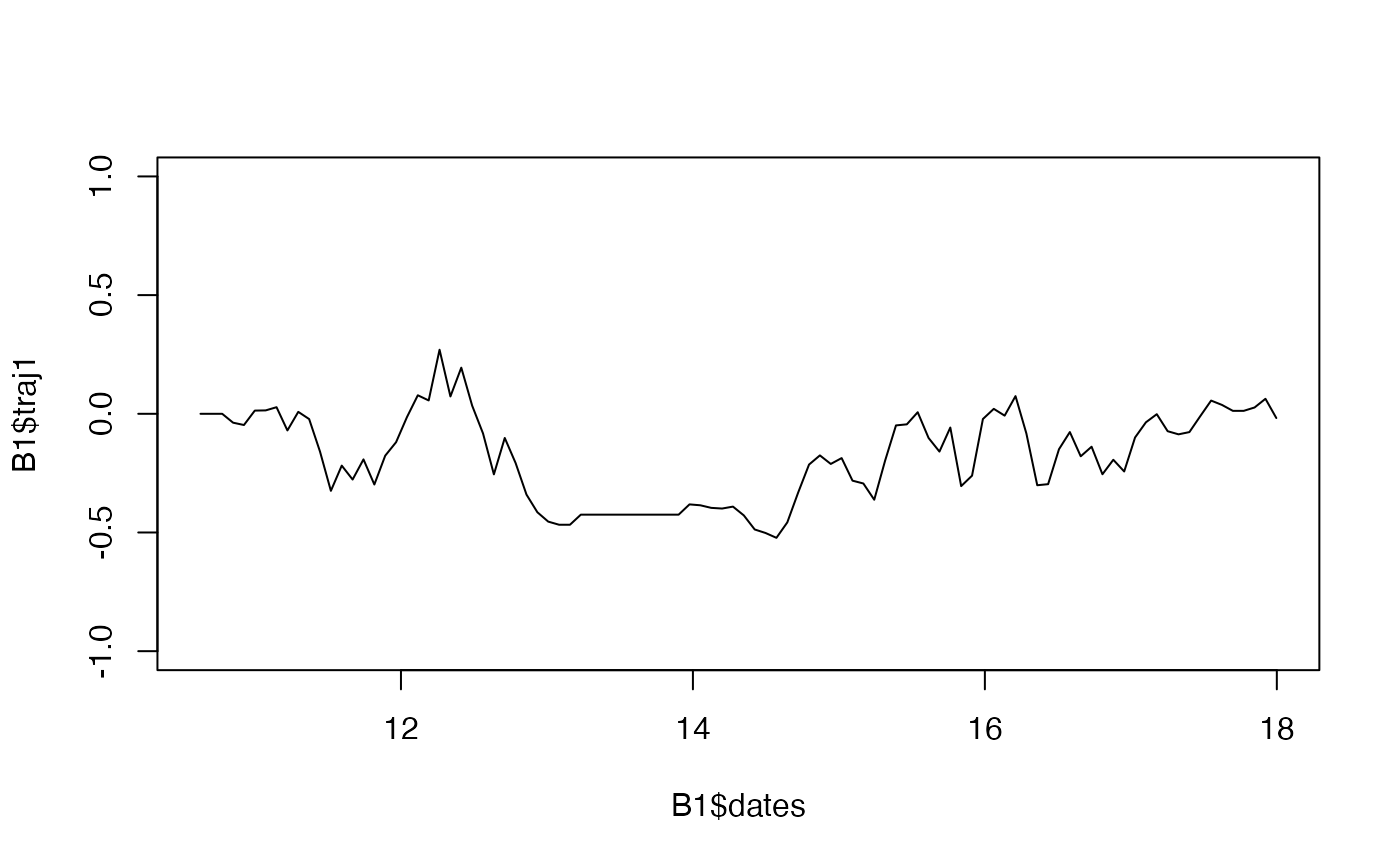
Simulate the trajectory of a Gaussian process, given a mean vector and a variance-covariance structure.
Arguments
- mean_vec
Vector (if multimensional) of means for the increments following gaussian distribution.
- varCov_mat
Corresponding variance-covariance structure.
- from
Initial time point at which the process is simulated.
- to
Last time point at which the process is simulated.
- start
Useful if the user wants to make the trajectory start from some given value.
- nb.points
Number of points at which the process is simulated.
Value
The trajectory of the Gaussian processes after simulating the multivariate Gaussian distributions with specified variance-covariance structure.
Author
Xavier Milhaud xavier.milhaud.research@gmail.com
Examples
list.comp <- list(f1 = "norm", g1 = "norm")
list.param <- list(f1 = list(mean = 12, sd = 0.4),
g1 = list(mean = 16, sd = 0.7))
sample1 <- rsimmix(n = 2000, unknownComp_weight = 0.5, comp.dist = list.comp,
comp.param = list.param)$mixt.data
## First get the variance-covariance matrix of the empirical process (Donsker correlation):
cov_mat <- .Call('_admix_estimVarCov_empProcess_Rcpp', PACKAGE = 'admix',
seq(from = min(sample1), to = max(sample1), length.out = 100), sample1)
## Plug it into the simulation of the gaussian process:
B1 <- sim_gaussianProcess(mean_vec=rep(0,nrow(cov_mat)), varCov_mat=cov_mat, from=min(sample1),
to = max(sample1), start = 0, nb.points = nrow(cov_mat))
plot(x = B1$dates, y = B1$traj1, type="l", xlim = c(min(sample1),max(sample1)), ylim = c(-1,1))